Directed Evolution
Methods for the Directed Evolution of Proteins
Proteins, the functional workhorses of living organisms, are composed of chains of interlinked amino acids that together form peptides and ultimately dictate the three-dimensional structure and function. Beneficial changes in the amino acid sequence and protein structure, likely a result from evolution and diverse natural selection pressures, is often a slow but necessary process that allows life to respond and adapt to environmental changes. Dramatically increasing the rate of change, directed evolution simulates the natural process to generate a library of proteins with diverse properties and occasionally enhanced functionality. Resulting in innovations in the fields of enzymology, drug development and medicine, the directed evolution of enzymes and the phage display of antibodies and peptides has been internationally recognized by awarding the pioneers in the field with the 2018 Nobel Prize in Chemistry.
Tuning protein functionalities using de novo and rational protein design
De novo protein design involves protein structural motif prediction and uses comparative modeling and fold recognition methods to arrive at the 3D structure.1 However, the predicted protein motifs derived from the amino acid sequence may not impart the same functionality to the target protein. In a rational design workflow, the sequence-structure-function dogma forms the central basis and involves identification of the target amino acid for mutation followed by site directed mutagenesis and other methods as outlined below. Following mutagenesis, the mutant proteins with altered functionalities are experimentally tested.2 The challenges imposed by rational design include low protein expression, stability, and achieving the desired functionality. It is also difficult to expand this approach to every single amino acid in a given protein. Moreover, these methods are difficult to attempt on proteins with no prior structural and or functional information.
Directed evolution and mimicking natural selection
Directed evolution is a robust method to design proteins with desirable functions. Unlike rational methods, directed evolution generates random mutations in the gene of interest and requires no protein structure information. Directed evolution mimics natural evolution by imposing stringent selection and screening methodologies to identify proteins with optimized functionality, including genetic diversity, binding, catalytic properties, thermal and environmental stability.3
History of directed evolution
With the generation of self-replicating RNA in 1967, directed evolution methodologies underwent exhaustive growth over next three decades. In the 1980s, methodologies for in vitro selection based on phage display were successfully employed to enrich the desired protein. In 1985, the discovery of PCR enabled the use of random and saturation mutagenesis methods as outlined in Table 1. Later, directed evolution methodologies improved functionality, enantioselectivity of many enzymes, and have been used widely to tailor metabolic pathways and various genomes.4,5
Directed evolution experimental design
A directed evolution experiment encompasses three essential steps namely, mutagenesis, screening, and gene amplification. Figure 1 provides an overview of the directed evolution methodology.
Generation of a mutant protein library
Defining the protein engineering goal for the desired functional diversification or substrate specificity is critical when generating a mutant protein library. Use of targeted libraries with a favorable ratio of beneficial over deleterious mutants is often an effective starting point. Table 1 summarizes the different methods used for mutagenesis and the principles behind them.6 The most common methods include error-prone PCR and DNA shuffling methods.
Selection and screening
After creating genetic diversification, the mutants are often transformed into bacterial or yeast hosts for protein expression and screened for functionality. Selection eliminates non-functional variants and is primarily based on plasmid, phage or a ribosome display, growth complementation and reporter-based strategies. During screening, individual protein variants are evaluated for desired activity. High-throughput methodologies enable effective screening for desired functionality and include the following tools and methods: direct microtiter plates, digital imaging coupled to spectroscopy, compartmentalization, FACS (fluorescence-activated cell sorting), cell surface display, resonance energy transfer, nuclear magnetic resonance (NMR), high-performance liquid chromatography (HPLC), gas chromatography and mass spectroscopy.7 The desired clones with the best functionality are identified after screening and serve as templates for the next round of gene manipulation. Systematic interplay by combining screening and selection methods will often result in the best variant with desired activity.
Gene amplification
The last step involves selecting the best mutant sequence or pool of sequences for amplification by PCR. Using the best sequence(s), the entire mutagenesis, screening, selection, and gene amplification cycle is repeated until a mutant containing the desired properties, as defined by the protein engineering goal, is identified.
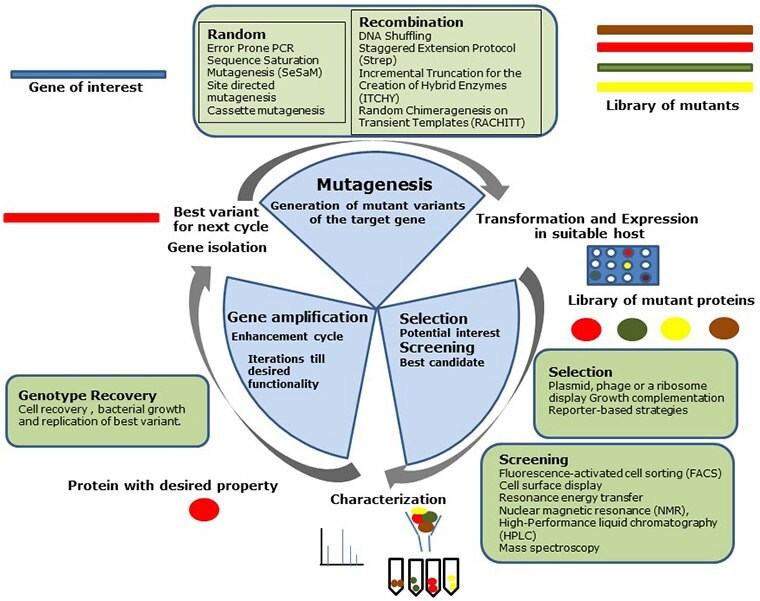
Figure 1: Steps in the directed evolution cycleSteps in the directed evolution cycle
Applications and successful protein engineering from utilizing directed evolution
Redesigning proteins by utilizing various mutagenesis methodologies has improved the functionality of many enzymes in terms of thermostability, specific activity, affinity, solubility, stability and enantioselectivity. Table 2 highlights some key enzymes that were identified using directed evolution.8
Success story of cytochrome P450 engineering
Cytochrome P450 enzymes are part of the drug metabolism pathway in mammals. It is a versatile enzyme with enormous substrate scope. Directed evolution of P450 enzymes has led to catalytic diversification and discovery of new biocatalytic transformations with unnatural P450 chemistry. The genetic tuning of P450 is an example of a biological change leading to the generation of new chemistry and expansion of the enzymes reaction space into a new biosynthetic pathway. The functionality of cytochrome P450, from Bacillus megaterium, was transformed from fatty acid hydroxylation to alkane degradation using directed evolution.9 Importantly, the integration of computational predictions coupled with directed evolution methodologies will likely expedite and strengthen our understanding on new protein functionality.10
Kapa Biosystems reagents from directed evolution methods
Directed evolution is a boon to life science and has advantages over natural evolution processes. Our suite of Kapa Biosystems reagents employs a directed evolution methodology that promotes natural selection in the lab to generate enzymes that are optimized for several life science applications, such as PCR, quantitative PCR (qPCR), next-generation sequencing and molecular diagnostics.
The PCR reagents that are available commercially consist of “wild-type” recombinant DNA polymerase (Taq polymerase) from Thermus aquaticus. However, our suite of Kapa Biosystems reagents consists of novel polymerase enzymes, synthesized using directed evolution technology, with enhanced specific activity, higher fidelity, increased processivity, and resistant to common PCR inhibitors. Kapa Biosystems provides a diverse range of reagents for PCR, qPCR and qRT-PCR kits for gene amplification.
Kapa Biosystems PCR and qPCR applications
Kapa Biosystems proteins, derived from directed evolution methodologies, contain numerous enhancements that are both diverse and unique compared to the wild-type proteins. The combination of enhanced enzymes from directed evolution methods coupled with additional factors, such as buffers and additional enzymes, are what make Kapa Biosystems kits so impactful. For example, KAPA Long Range PCR Kits contain a Taq DNA polymerase and a modified archaeal (B-family) DNA polymerase with proofreading activity. KAPA Long Range HotStart DNA Polymerase has been used in the detection of hybrid genes by Long-Range PCR,11 while KAPA2G Fast Multiplex PCR Kits (see Figure 2), with significantly faster extension times than wild-type Taq, have been used to analyze three single nucleotide polymorphisms (SNP's; rs1049673, rs3211931, and rs3212162) in Alzheimer's patients by use of multiplexed PCR amplification, followed by a single base extension.12
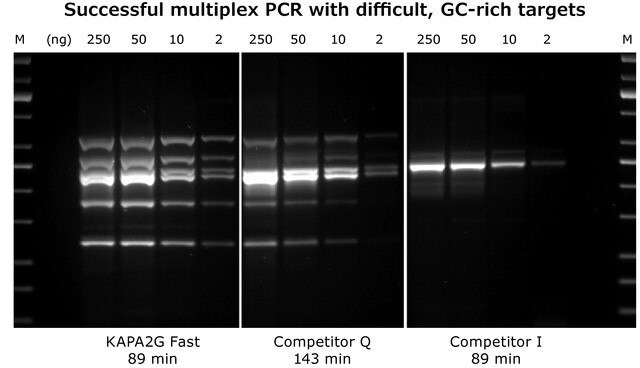
Figure 2: KAPA2G Fast Multiplex PCR Kit: 6-plex multiplex PCR with the KAPA2G Fast Multiplex PCR Kit, Competitor Q and Competitor I using the same cycling conditions (30 cycles).
Kapa Biosystems qPCR reagents and kits have been optimized to identify low copy and difficult targets with improved reproducibility. In order to analyze crude samples, KAPA PROBE FORCE qPCR Kits contain a master mix that removes the need for DNA purification and is resistant to inhibitors from blood, tissue and plant samples. Additionally, the KAPA PROBE FAST qPCR Kit (Figure 3) is suitable for extremely sensitive and accurate real-time PCR using sequence-specific fluorogenic probe chemistries, such as hydrolysis probes, FRET probes, and displacement probes. KAPA PROBE FAST qPCR has been used to determine the expression level of SOX7 mRNA and other developmentally important factors.13
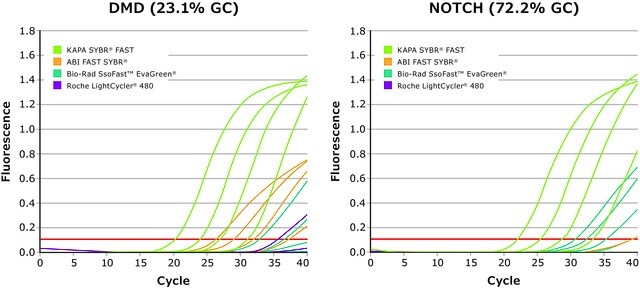
Figure 3: KAPA SYBR® FAST Kit: Superior performance and quantification for KAPA SYBR® FAST compared to competitor kits for DMD and NOTCH target gene amplification using human genomic DNA.
Summary
Directed evolution is a powerful strategy for improving the functionality of proteins. Generating thousands of variants, directed evolution uses high throughput screening to arrive at the best solution when either the biological function is obscure or when chemical knowledge on the substrate and protein structure is limited. Directed evolution has enabled the generation of numerous engineered proteins, including pure isomers of pharmaceuticals and proteins for critical life-science applications. Provided the rapid turnaround time for generating new proteins through directed evolution, engineered proteins and enzymes will likely continue to have a significant impact in advancing our understanding of numerous biological processes and biocatalysis.14
References
To continue reading please sign in or create an account.
Don't Have An Account?