P4929
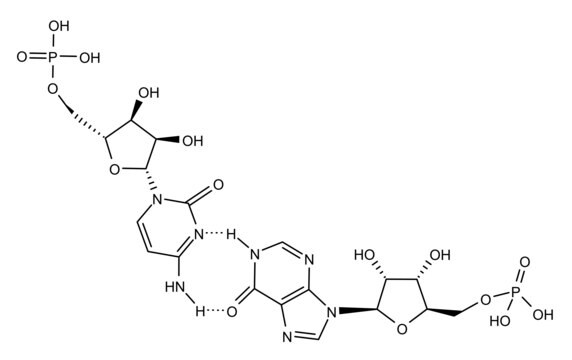
Poly(deoxyinosinic-deoxycytidylic) acid sodium salt
double-stranded alternating copolymer
Synonym(s):
Poly(dI-dC) • Poly(dI-dC) sodium salt
Sign Into View Organizational & Contract Pricing
All Photos(1)
About This Item
Recommended Products
Looking for similar products? Visit Product Comparison Guide
Application
Poly(deoxyinosinic-deoxycytidylic) acid (Poly(dI-dC) • Poly(dI-dC)) is an alternating copolymer used as a DNA substrate for evaluation of DNA methytransfeases, such as DNA-methyltransferase 1 and as double-stranded DNA model for conformational studies of DNA structure dynamics and drug, small molecule, interactions.
Poly(deoxyinosinic-deoxycytidylic) acid sodium salt has been used in a study that synthesized and characterized bis(2-(pyrimidin-2-γl)ethoxy)alkanes and their pharmacological activity. Poly(deoxyinosinic-deoxycytidylic) acid sodium salt has also been used in a study that investigated the growth of calcium carbonate films on LB/LbL matrices.
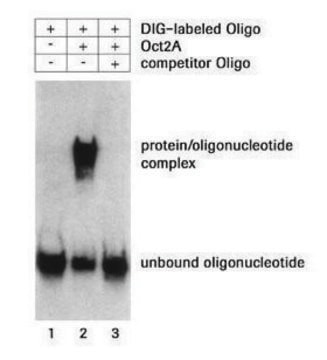
Poly(deoxyinosinic-deoxycytidylic) acid sodium salt has been used in the electrophoretic mobility shift assay in p65 protein and neuro-2A protein lysate. It has also been used as a substrate in the protein arginine methyltransferase 1 (PRMT1) and DNA methyltransferase 1 (DNMT1) selectivity assay.
Biochem/physiol Actions
Poly(2′-deoxyinosinic-2′-deoxycytidylic acid), poly(dI-dC) is a synthetic DNA substrate. At high salt, Poly(dI-dC) exists in left handed helical conformation and reverts to right-handed form upon decreasing salt. It is used in electrophoretic mobility shift assay (EMSA).
Other Notes
Double-stranded alternating copolymer
Unit Definition
One unit will yield an A260 of 1.0 in 1.0 ml of 20 mM sodium phosphate/100 mM NaCl, pH 7.0 (1 cm light path)
Storage Class
11 - Combustible Solids
wgk_germany
WGK 3
flash_point_f
Not applicable
flash_point_c
Not applicable
ppe
Eyeshields, Gloves, type N95 (US)
Choose from one of the most recent versions:
Certificates of Analysis (COA)
Lot/Batch Number
Don't see the Right Version?
If you require a particular version, you can look up a specific certificate by the Lot or Batch number.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Customers Also Viewed
Identification of Selective, Cell Active Inhibitors of Protein Arginine Methyltransferase 5 through Structure-Based Virtual Screening and Biological Assays
Ye F, et al.
Journal of Chemical Information and Modeling, 58(5), 1066-1073 (2018)
Dimitrios Priftis et al.
Langmuir : the ACS journal of surfaces and colloids, 28(23), 8721-8729 (2012-05-15)
A systematic study of the interfacial energy (γ) of polypeptide complex coacervates in aqueous solution was performed using a surface forces apparatus (SFA). Poly(L-lysine hydrochloride) (PLys) and poly(L-glutamic acid sodium salt) (PGA) were investigated as a model pair of oppositely
Meaghan L Clark et al.
Inorganic chemistry, 47(20), 9410-9418 (2008-09-25)
This paper focuses on DNA-binding interactions exhibited by Pt(dma-T)CN(+), where dma-T denotes 4'-dimethylamino-2,2':6',2''-terpyridine, and includes complementary studies of the corresponding pyrr-T complex, where pyrr-T denotes 4'-(N-pyrrolidinyl)-2,2':6',2''-terpyridine. The chromophores are useful for understanding the interesting and rather intricate DNA-binding interactions exhibited
Dexamethasone treatment of calves latently infected with bovine herpesvirus 1 leads to activation of the bICP0 early promoter, in part by the cellular transcription factor C/EBP-alpha
Workman A, et al.
Journal of Virology, 83(17), 8800-8809 (2009)
Ligang Fan et al.
Nature communications, 11(1), 4947-4947 (2020-10-04)
Pseudomonas syringae is a Gram-negative and model pathogenic bacterium that causes plant diseases worldwide. Here, we set out to identify binding motifs for all 301 annotated transcription factors (TFs) of P. syringae using HT-SELEX. We successfully identify binding motifs for
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service

![Poly[d(I-C)] lyophilized, pkg of 10 U (10108812001 [A<sub>260</sub> units]), pkg of 50 U (11219847001 [A<sub>260</sub> units])](/deepweb/assets/sigmaaldrich/product/images/352/091/ef743cea-ccd8-44f1-8f3b-dec5a1e4f5d1/640/ef743cea-ccd8-44f1-8f3b-dec5a1e4f5d1.jpg)