Differentiation and ID Media for Salmonella
Today’s use of classical culture methods to identify Salmonella, a potent food-borne pathogen
Salmonella contamination is the second leading cause of food-borne illness worldwide. Controlling outbreaks of Salmonella is an important task for food regulators, restaurants and the food industry in general.
The Salmonella family includes over 2,300 serotypes of bacteria, but two types, Salmonella enteritidis and Salmonella typhimurium, are responsible for about half of all human infections. Most outbreaks of Salmonella are traced back to dairy, poultry and meat products, but Salmonella can grow on nearly any food. Chicken, eggs and their derivative products are particularly high risk.
Figure 1. Salmonella Bacteria
Microbiological control in the food industry plays a critical role in preventing Salmonella outbreaks. Tests and media used for identification of Salmonella take advantage of unique aspects of Salmonella physiology or biochemistry relative to other genera within the family Enterobacteriaceae. For example, bacteria from the genus Salmonella are mostly facultative anaerobes, oxidase-negative, catalase-positive and gramnegative rods. Most strains are motile and ferment glucose with production of both acid and gas. The media used currently for the differentiation and identification of Salmonella are still based on the detection of carbohydrate fermentation indicated by a pH indicator (see also Table 1 for carbohydrate fermentation ability), the detection of proteolytic activity, hydrogen sulphide production and selectivity. Most modern media also combine some of this detection system to make the media more reliable. A listing of the most common selective and differential media appears in Table 2.
| Fermentation | Cat. No. | ||
|---|---|---|---|
| Carbohydrate | Acid | Gas | Cabohydrate Discs |
| Adonitol | - | - | 55876 |
| Arabinose | +/- | +/- | 80372 |
| Cellobiose | - | - | 56481 |
| Dextrose | + | +/- | 63367 |
| Dulcitol | +/- | +/- | 73044 |
| Fructose | +/- | +/- | 53901 |
| Galactose | + | +/- | 89608 |
| Inositol | +/- | +/- | 89614 |
| Lactose | - | - | 28816 |
| Maltose | + | +/- | 77653 |
| Mannitol | + | +/- | 94438 |
| Mannose | +/- | +/- | 94445 |
| Melibiose | + | + | 93196 |
| Raffinose | - | - | 94226 |
| Rhamnose | +/- | +/- | 93999 |
| Salicin | - | - | 92971 |
| Sorbitol | + | +/- | 93998 |
| Sucrose | - | - | 94309 |
| Trehalose | + | +/- | 92961 |
| Xylose | + | +/- | 07411 |
| Brand | Cat # | Name |
|---|---|---|
| Sigma-Aldrich | A0715 | Andrade Peptone Water |
| Sigma-Aldrich | 28943 | Andrade peptone water, Vegitone |
| Sigma-Aldrich | 95388 | Bismuth sulfite Agar |
| Sigma-Aldrich | 15835 | BPL Agar |
| Sigma-Aldrich | 70134 | Brilliant Green Agar, modified |
| Sigma-Aldrich | 16026 | Brilliant Green Phenol Red Lactose Sucrose Agar |
| Sigma-Aldrich | 36408 | Bromcresol Purple Broth |
| Sigma-Aldrich | 22520 | China Blue Lactose Agar |
| Sigma-Aldrich | 55420 | CLED Agar |
| Sigma-Aldrich | 70135 | DCLS Agar |
| Sigma-Aldrich | 90035 | DCLS Agar No. 2 |
| Sigma-Aldrich | D2935 | Decarboxylase Broth Base, Moeller |
| Sigma-Aldrich | D7809 | Deoxycholate Citrate Agar |
| Sigma-Aldrich | E5399 | Endo Agar |
| Sigma-Aldrich | 70137 | ENDO Agar (Base) |
| Sigma-Aldrich | 16447 | Glucose Bromcresol Purple Agar |
| Sigma-Aldrich | 51490 | Hektoen Enteric Agar |
| Sigma-Aldrich | 60787 | Kligler Agar |
| Sigma-Aldrich | 61792 | Leifson Agar |
| Sigma-Aldrich | 66304 | Lysine Decarboxylase Broth |
| Sigma-Aldrich | 62915 | Lysine Iron Agar |
| Sigma-Aldrich | 70143 | Mac Conkey Agar No 1 |
| Sigma-Aldrich | 19352 | Mac Conkey Agar No 1, Vegitone |
| Sigma-Aldrich | M8302, 94216 | MacConkey Agar with Crystal Violet, Sodium Chloride and 0.15% Bile Salts |
| Sigma-Aldrich | 70144 | MacConkey Broth |
| Sigma-Aldrich | 75717, 16377 | MacConkey broth purple |
| Sigma-Aldrich | 63014 | MacConkey MUG Agar |
| Sigma-Aldrich | 51405 | MacConkey-Agar (without salt) |
| Sigma-Aldrich | 69965 | Mossel Broth |
| Sigma-Aldrich | 43052 | Muller-Kauffmann Tetrathionate Broth, Base (ISO) |
| Sigma-Aldrich | 75315 | OF Test Nutrient Agar |
| Sigma-Aldrich | 81648 | Pril® Mannitol Agar |
| Sigma-Aldrich | 04584 | Rappaport Vassiliadis Broth acc. to DIN EN ISO 6579:2002 |
| Sigma-Aldrich | 17173 | Rappaport Vassiliadis Broth, modified |
| Sigma-Aldrich | R0773 | Rappaport Vassiliadis Medium |
| Sigma-Aldrich | 92322 | Rappaport Vassiliadis medium (base), modified, semi-solid |
| Sigma-Aldrich | 84368 | Salmonella Agar according to Önöz |
| Sigma-Aldrich | 84370 | Salmonella Enrichment Broth |
| Sigma-Aldrich | 70153 | Selenite Broth (Base) |
| Sigma-Aldrich | 84922 | Selenite Cystine Broth |
| Sigma-Aldrich | 85438 | SIM Medium |
| Sigma-Aldrich | 85463 | Simmons Citrate Agar |
| Sigma-Aldrich | 85640 | SS-Agar |
| Sigma-Aldrich | 86352 | TBG Broth |
| Sigma-Aldrich | 88151 | Tetrathionate Broth |
| Sigma-Aldrich | 88148 | Tetrathionate Enrichment Broth according to Muller-Kauffmann |
| Sigma-Aldrich | 44940 | Triple Sugar Iron Agar |
| Sigma-Aldrich | 51463 | Urea Broth |
| Sigma-Aldrich | 42376 | Violet Red Bile Agar, Vegitone |
| Sigma-Aldrich | 70189, 79873 | Violet Red Bile Glucose Agar |
| Sigma-Aldrich | 17213 | Violet Red Bile Glucose Agar without Lactose |
| Sigma-Aldrich | 53605 | Violet Red Bile Glucose Agar without Lactose, Vegitone |
| Sigma-Aldrich | 41270 | Violet Red Bile Lactose Dextrose Agar |
| Sigma-Aldrich | 95273 | VRB MUG Agar |
| Sigma-Aldrich | 95586 | XLD Agar |
| Sigma-Aldrich | 76721 | XLT4 Agar (Base) |
In addition, our current technology offers the chromogenic media, which makes identification even more reliable and faster as they detect a characteristic enzyme of the Salmonella. These reactions are based on the cleavage of a chromogenic substrate which results in a visible color change (see Table 3).
| Brand | Cat # | Name |
|---|---|---|
| Sigma-Aldrich | 00563 | HiCrome™ MM Agar |
| Sigma-Aldrich | 90918 | HiCrome™ RajHans Medium, Modified |
| Sigma-Aldrich | 78419 | HiCrome™ Salmonella Agar |
| Sigma-Aldrich | 05538 | HiCrome™ Salmonella Agar, Improved |
| Sigma-Aldrich | 84369 | Salmonella Chromogen Agar |
| Sigma-Aldrich | 01993 | Salmonella Chromogen Agar Set |

Figure 2.HiCrome™ Salmonella Agar, Improved
Detection of hydrogen sulfide Production of microorganisms such as Salmonella
A large number of bacteria can produce H2S in small amounts from sulfur containing amino acids in carbohydrate media. When combined with lead acetate, the H2S will produce a black precipitate, giving rise to a visible black coloured reaction on the paper strip. The lead acetate method is very sensitive, allowing the detection of trace levels of hydrogen sulphide.
Test with strips: Inoculate peptone water (Cat. No. 70179) with the suspect organism. Insert a lead acetate paper strip between the plug and inner wall of tube, above the inoculated medium and incubate at 35 °C for 18-24 hours. A positive reaction appears as a blackening of the lower part of the strip. In the case of negative response, no blackening should appear (see Figure 3).
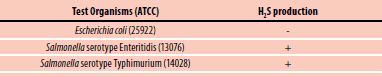
Figure 2.1. Test OrganismsTest Organisms

Figure 3.Hydrogen Sulfide Test Strips
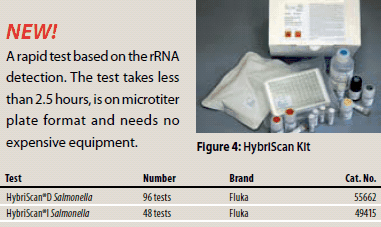
Figure 4.A rapid test based on the rRNA detection
To continue reading please sign in or create an account.
Don't Have An Account?